Bioinformatics and systems biology approaches identify the potential common pathogenesis of periodontitis and rheumatoid arthritis
##plugins.themes.bootstrap3.article.main##
Abstract
Abstract
Rheumatoid arthritis (RA) and periodontitis (PD) are chronic inflammatory disorders with significant comorbidity, yet their shared pathogenic mechanisms remain poorly understood.
Methods
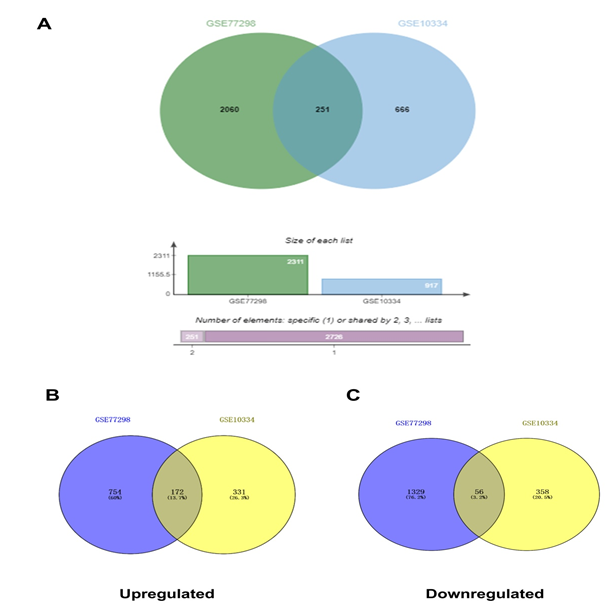
This study employed bioinformatics approaches to identify common molecular pathways and diagnostic biomarkers linking RA and PD. Datasets for RA and PD were obtained from the GEO database. Common differentially expressed genes (DEGs) were identified, followed by gene and pathway enrichment analysis, protein-protein interaction (PPI) network construction, hub gene identification, and immune infiltration analysis. Interaction networks involving transcription factor (TF)-gene, gene-miRNA, gene-disease, and protein-drug were established based on the identified hub genes.Diagnostic accuracy of hub genes was evaluated using receiver operating characteristic (ROC) curve analysis, and expression profiles were validated using external datasets.
Results
A total of 251 common DEGs were identified, with 15 hub genes (PTPRC, ITGB2, CXCL8, CCL5, CXCR4, MMP9, FCGR3B, CD2, SELL, CXCL1, LYN, FCER1G, CXCL2, IL7R, and CD69) showing high diagnostic performance (AUC > 0.7) across independent datasets. Immune infiltration analysis revealed significant correlations between hub genes and immune cells, particularly B cells and CD4+ T cells. TFs, miRNAs, and potential therapeutic drugs associated with hub genes were also predicted.
Conclusion
This study identified shared molecular mechanisms between RA and PD, highlighting potential diagnostic biomarkers and therapeutic targets. The findings provide a foundation for future research into comorbid disease management.
Keywords: periodontitis, rheumatoid arthritis, differentially expressed genes, bioinformatics, hub genes