Study on the Bacterial Community Structure of Sediment in Different Habitats of Penaeus Orientalis
##plugins.themes.bootstrap3.article.main##
Abstract
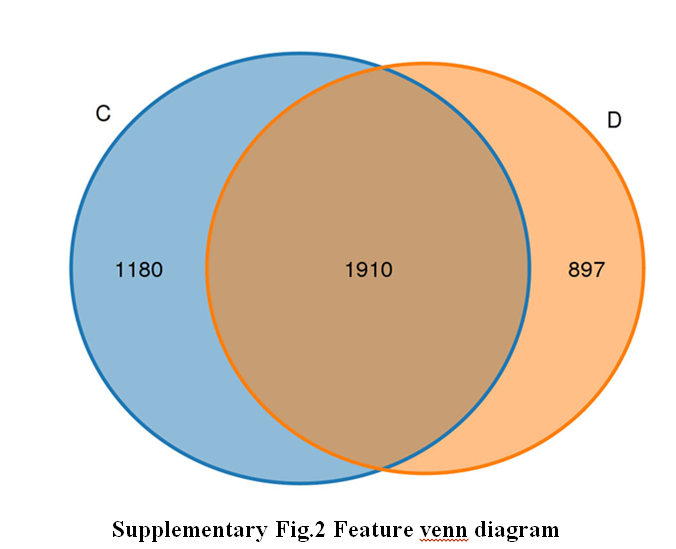
The microbial communities in Penaeus orientalis' habitats are important to the quality of the species. This study utilized the PacBio sequencing platform to analyze the microbial diversity in the different habitats of the species. The analysis indicates that the shrimp pond sediment samples have a greater number of features compared to the intertidal gully sediment samples, with a total of 1910 feature sequences overlapping between the two. According to the results of beta diversity analysis showed that the sediment samples in the intertidal gully were far away from the sediment samples in the shrimp pond, indicating a great difference in microbial flora species. Sediment samples from shrimp ponds showed the presence of dominant bacteria genera such as Sulfurovum and Lactobacillus. According to KEGG functional prediction analysis, the relative abundance of the Translation metabolic pathway in sediment samples was significantly higher compared to that of intertidal gullies sediment samples. In both samples, the highest abundance of metabolic pathways was Translation, whereas the abundance of metabolic pathways related to Signaling molecules and interaction metabolic pathways was relatively low. The COG function prediction analysis revealed that the abundance of Signal transduction mechanisms in the sediment samples from intertidal gullies was higher than those in the samples from shrimp pools. The results of this study analyzed the environmental adaptability of P. orientalis from the perspective of habitat environment and provided a basis for the development of strains that have special functions and natural metabolites in the sediment.