Identification and Functional Prediction of Lncrnas Associated with Oocyte Development in the Donkey
##plugins.themes.bootstrap3.article.main##
Abstract
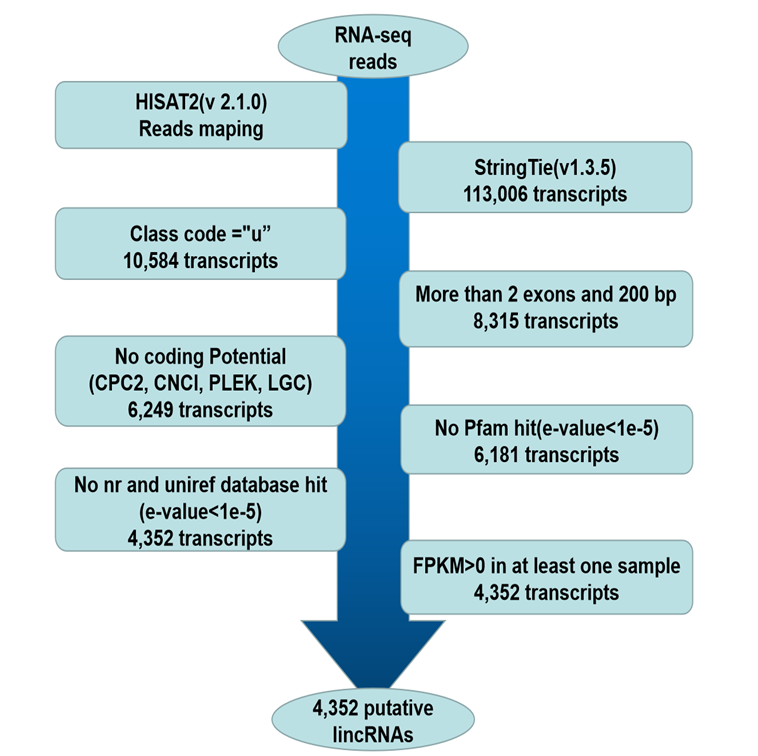
Long non-coding RNAs (lncRNAs) play key regulatory roles in various biological processes. However, the importance and molecular regulatory mechanisms of lncRNAs in donkey oocyte maturation remain to be further investigated. This study used published transcriptomic data from the germinal vesicle (GV) phase and metaphase II (MII) phase donkey oocytes to identify lncRNAs and obtained 4,352 novel lncRNAs. Compared with the coding genes, the novel lncRNAs and the known lncRNAs exhibited some typical features, such as shorter transcript length and fewer exons. A total of 3,805 coding genes and 569 lncRNAs were differentially expressed between the GV and MII stages. The differentially expressed genes were found to be involved in various biological processes related to oocyte development. The potential target genes of differentially expressed lncRNAs were predicted in cis and trans manner. Functional analysis of lncRNA targets showed that some lncRNAs may act on potential target genes involved in oocyte maturation. This study provides valuable information for further investigation of the molecular mechanisms of oocyte development in donkeys, serving as a theoretical basis for improving the in vitro maturation rate and ultimately benefiting from the expansion of the donkey population and the conservation of biodiversity and genetic resources.